GENETIC MAPPING III. The problem of double crossovers in genetic
By A Mystery Man Writer
Last updated 20 Sept 2024
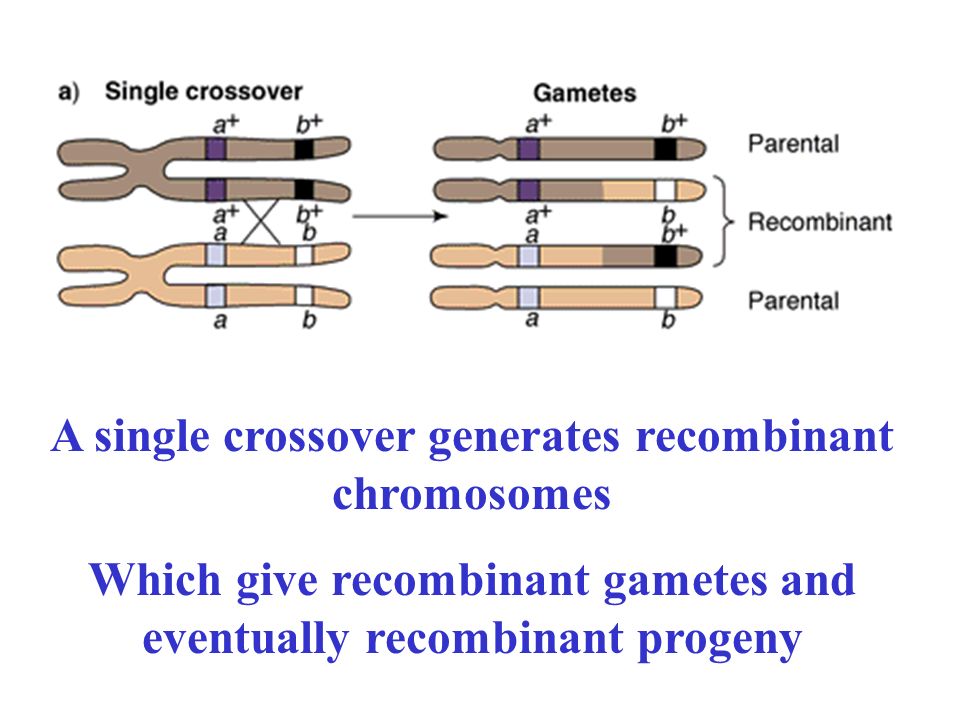
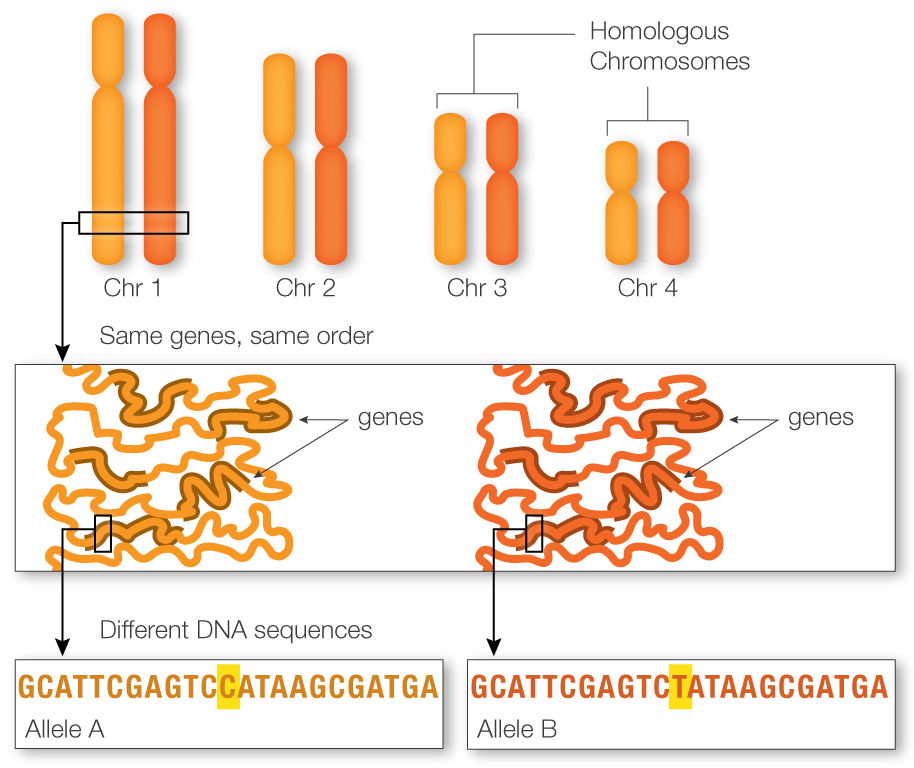
Consider a cross to map 2 genes, a and b They are some distance apart, but mappable The heterozygote is in tetrad stage: a+ b+/ab
The problem of double crossovers in genetic mapping experiments
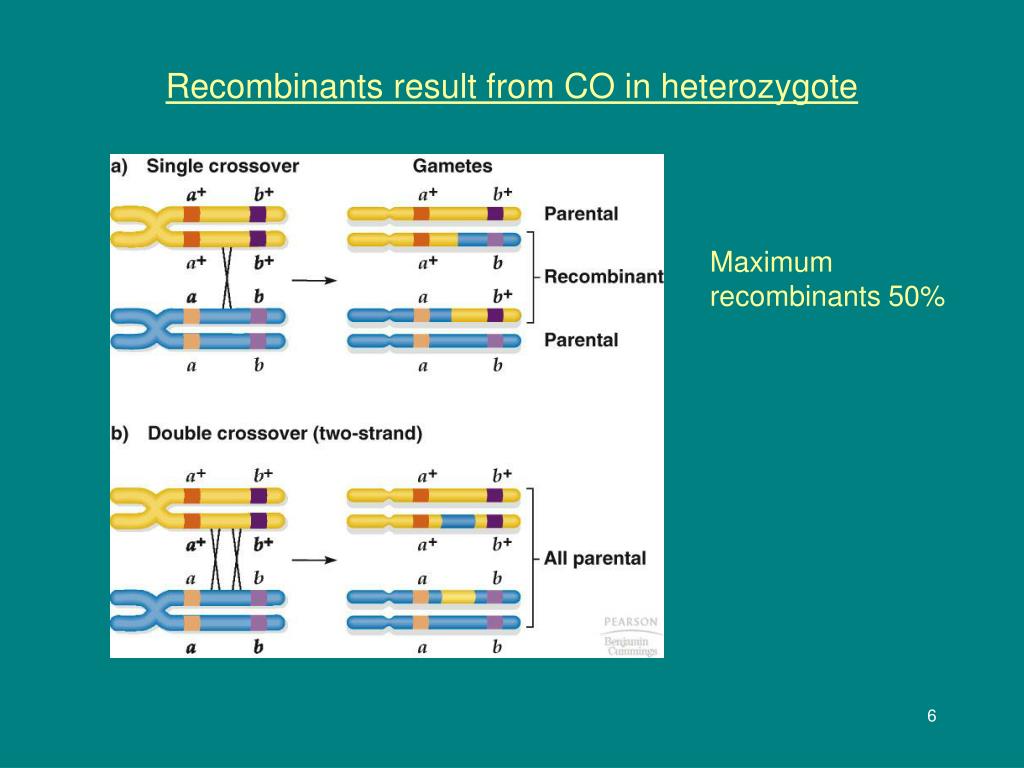
A single crossover generates recombinant chromosomes Which give recombinant gametes and eventually recombinant progeny
A 2-strand, double crossover restores the original arrangement of the marker genes So all progeny are scored as parental, with no recombinants
It looks exactly as if there has been no crossing over There have been two crossover events which will be uncounted
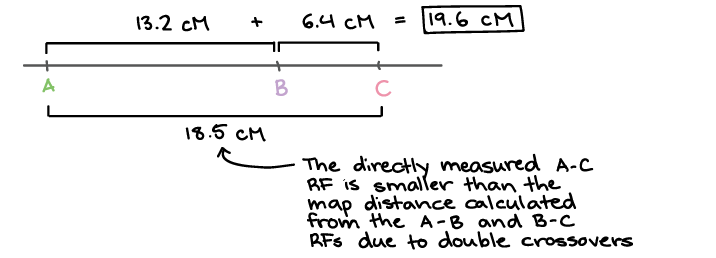
And since recombination frequency is a measure of map distance, this means that the distance between the genes will be underestimated
How can we avoid these errors Two general ways:
1. Map closely linked genes Double crossovers rarely occur within map distances < 10 cM
Do three-point testcrosses, rather than two-point These involve 3 genes within a relatively short section of chromosome.
The rationale for using these is illustrated in the next slide.
As before, a 2-strand double crossover gives gametes that are nonrecombinant for genes a and b
BUT, notice that the resulting gametes are recombinant with respect to c
The gene in the middle reveals the occurrence of a double crossover 3-point crossovers are routinely used for mapping, because they allow us to correct for double crossovers, and determine the gene order
Suppose we want to map 3 genes in a plant Fruit color: p = purple; p+ = yellow Fruit shape: r = round; r+ = elongated Juiciness: j = juicy; j+ = dry
What is the order, and map distances, of these 3 genes
We set up our testcross with a triply heterozygous parent, in coupling phase (in this case) and count the offspring
We know that is the genes were unlinked, we would expect eight phenotypic classes of progeny
For this kind of trihybrid cross, we expect the same classes, but not in the same proportions
Because of linkage, some phenotypic classes may have 0 individuals; if so, that’s important to note
Here are the eight phenotypic classes of progeny
These are parentals. Note that they are in approximately equal numbers
These recombinants both involve the p gene Notice that they are in about equal numbers, and are rarer than the parentals
These recombinants involve the r gene They are rarer still
These recombinants are the rarest. Gene j is involved
We expect double crossovers to be rarer than single crossovers So it follows that recombinants due to double crossovers will be the rarest class
We can use this fact to help us order the genes. How Recall our earlier example:
Notice how the double crossover restored the outside genes to the parental arrangement, but the middle gene has its orientation changed
So the gene which is in a recombinant arrangement in the rarest, double crossover class of progeny, must be the middle gene.
We can see that p and r are in their parental configuration, but j is in a new arrangement So, j must be the gene in the middle The order must be p, j, r
Now that we know the correct gene order, we can interpret the data to generate map distances:
For the p - j distance, we need to add together all the recombinant progeny resulting from crossovers in Region I
This includes both single crossovers and the double crossovers (since they also involve this region of the chromosome)
So, the percentage of recombinants = [(52+46) + (4+2)]/500 x 100% = 104/500 x 100% = 20.8% So, p and j are 20.8 cM apart
We do the same sort of calculations to find the distance between j and r
We again add together the single crossovers (this time from Region II) and the double crossovers
[(22+22) + (4+2)]/500 x 100% = 50/500 x 100% = 10.0% So, j and r are 10.0 cM apart
To get the distance between p and r, we simply add the inner distances = 30.8 cM.
The problem of double crossovers in genetic mapping experiments
A single crossover generates recombinant chromosomes Which give recombinant gametes and eventually recombinant progeny
A 2-strand, double crossover restores the original arrangement of the marker genes So all progeny are scored as parental, with no recombinants
It looks exactly as if there has been no crossing over There have been two crossover events which will be uncounted
And since recombination frequency is a measure of map distance, this means that the distance between the genes will be underestimated
How can we avoid these errors Two general ways:
1. Map closely linked genes Double crossovers rarely occur within map distances < 10 cM
Do three-point testcrosses, rather than two-point These involve 3 genes within a relatively short section of chromosome.
The rationale for using these is illustrated in the next slide.
As before, a 2-strand double crossover gives gametes that are nonrecombinant for genes a and b
BUT, notice that the resulting gametes are recombinant with respect to c
The gene in the middle reveals the occurrence of a double crossover 3-point crossovers are routinely used for mapping, because they allow us to correct for double crossovers, and determine the gene order
Suppose we want to map 3 genes in a plant Fruit color: p = purple; p+ = yellow Fruit shape: r = round; r+ = elongated Juiciness: j = juicy; j+ = dry
What is the order, and map distances, of these 3 genes
We set up our testcross with a triply heterozygous parent, in coupling phase (in this case) and count the offspring
We know that is the genes were unlinked, we would expect eight phenotypic classes of progeny
For this kind of trihybrid cross, we expect the same classes, but not in the same proportions
Because of linkage, some phenotypic classes may have 0 individuals; if so, that’s important to note
Here are the eight phenotypic classes of progeny
These are parentals. Note that they are in approximately equal numbers
These recombinants both involve the p gene Notice that they are in about equal numbers, and are rarer than the parentals
These recombinants involve the r gene They are rarer still
These recombinants are the rarest. Gene j is involved
We expect double crossovers to be rarer than single crossovers So it follows that recombinants due to double crossovers will be the rarest class
We can use this fact to help us order the genes. How Recall our earlier example:
Notice how the double crossover restored the outside genes to the parental arrangement, but the middle gene has its orientation changed
So the gene which is in a recombinant arrangement in the rarest, double crossover class of progeny, must be the middle gene.
We can see that p and r are in their parental configuration, but j is in a new arrangement So, j must be the gene in the middle The order must be p, j, r
Now that we know the correct gene order, we can interpret the data to generate map distances:
For the p - j distance, we need to add together all the recombinant progeny resulting from crossovers in Region I
This includes both single crossovers and the double crossovers (since they also involve this region of the chromosome)
So, the percentage of recombinants = [(52+46) + (4+2)]/500 x 100% = 104/500 x 100% = 20.8% So, p and j are 20.8 cM apart
We do the same sort of calculations to find the distance between j and r
We again add together the single crossovers (this time from Region II) and the double crossovers
[(22+22) + (4+2)]/500 x 100% = 50/500 x 100% = 10.0% So, j and r are 10.0 cM apart
To get the distance between p and r, we simply add the inner distances = 30.8 cM.
Chapter 7 Flashcards
Gene mapping by a Three-Point Test-Cross - ppt download
PPT - Genomics Lecture Topics Genetic mapping studies: two
Genetic Linkage
PPT - Integrated Physical and Genetic Mapping of Upland Cotton
The Inheritance of Spore Color - ppt download
PPT - Geneomics and Database Mining and Genetic Mapping PowerPoint
Genetic linkage & mapping (article)
Solved BIO340 Activity #4: 3-Point Cross, Gene Order and
PPT - Genetic Mapping in a Three-Point Test Cross PowerPoint
PPT - Chapters 14 - Genetic Mapping in Eukaryotes : Discovery of
PPT - Genetic Mapping PowerPoint Presentation, free download - ID
Recommended for you
 Double Cross Vodka14 Jul 2023
Double Cross Vodka14 Jul 2023 Figure S3. Double crossover (CO) observed in genetic interval with14 Jul 2023
Figure S3. Double crossover (CO) observed in genetic interval with14 Jul 2023 Genetic Mapping: Three-point Testcross and Double Crossover14 Jul 2023
Genetic Mapping: Three-point Testcross and Double Crossover14 Jul 2023 Genetics chapter 5 part 2(1)14 Jul 2023
Genetics chapter 5 part 2(1)14 Jul 2023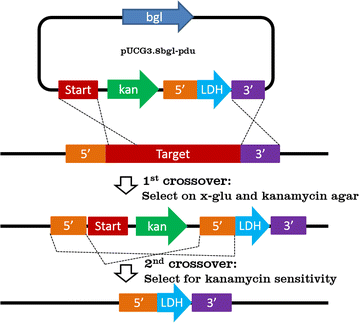 Development of an efficient technique for gene deletion and14 Jul 2023
Development of an efficient technique for gene deletion and14 Jul 2023 Schematic depiction of the single-step double recombination14 Jul 2023
Schematic depiction of the single-step double recombination14 Jul 2023 Drivers react positively to double crossover diamond interchange14 Jul 2023
Drivers react positively to double crossover diamond interchange14 Jul 2023 Genetics 03: 'Linkage and tetrad analysis in yeast14 Jul 2023
Genetics 03: 'Linkage and tetrad analysis in yeast14 Jul 2023 All Types of Cross Over Railway Turnouts for Rail Track System14 Jul 2023
All Types of Cross Over Railway Turnouts for Rail Track System14 Jul 2023- double crossover - Wiktionary, the free dictionary14 Jul 2023
You may also like
 Barra de balé portátil Modelo 3D - TurboSquid 200730414 Jul 2023
Barra de balé portátil Modelo 3D - TurboSquid 200730414 Jul 2023 Vanity Fair Illumination Front Close Full Coverage Underwire Bra 75339 Midnight Black14 Jul 2023
Vanity Fair Illumination Front Close Full Coverage Underwire Bra 75339 Midnight Black14 Jul 2023 Seamless Sports Bra with 10% discount!14 Jul 2023
Seamless Sports Bra with 10% discount!14 Jul 2023 Workout Routines & Training Programs - Muscle & Fitness14 Jul 2023
Workout Routines & Training Programs - Muscle & Fitness14 Jul 2023 Oroblu Shock Up Shaping Pantyhose14 Jul 2023
Oroblu Shock Up Shaping Pantyhose14 Jul 2023 Sexy Design Lovely Super Slim Leggings Heart Pattern Pantyhose Stylish – the best products in the Joom Geek online store14 Jul 2023
Sexy Design Lovely Super Slim Leggings Heart Pattern Pantyhose Stylish – the best products in the Joom Geek online store14 Jul 2023 FRALDA PERSONAL SOFT E PROTECT É BOA, USANDO DURANTE O DIA POR 6H14 Jul 2023
FRALDA PERSONAL SOFT E PROTECT É BOA, USANDO DURANTE O DIA POR 6H14 Jul 2023 Baby Blue Background With Small White Flowers And Bokeh With Copy14 Jul 2023
Baby Blue Background With Small White Flowers And Bokeh With Copy14 Jul 2023 Yo Gabba Gabba! on X: DJ Lance Rock, Muno & Plex @Moogfest14 Jul 2023
Yo Gabba Gabba! on X: DJ Lance Rock, Muno & Plex @Moogfest14 Jul 2023 The Difference between Lace Closure and Lace Frontal, and How to Choose?14 Jul 2023
The Difference between Lace Closure and Lace Frontal, and How to Choose?14 Jul 2023